OntoPortal Alliance
Dedicated to promoting semantic services.
Our mission
The OntoPortal Alliance is a consortium of several research and infrastructure teams and a company dedicated to promoting the development of ontology repositories ––in science and other disciplines–– based on the open, collaboratively developed OntoPortal software.
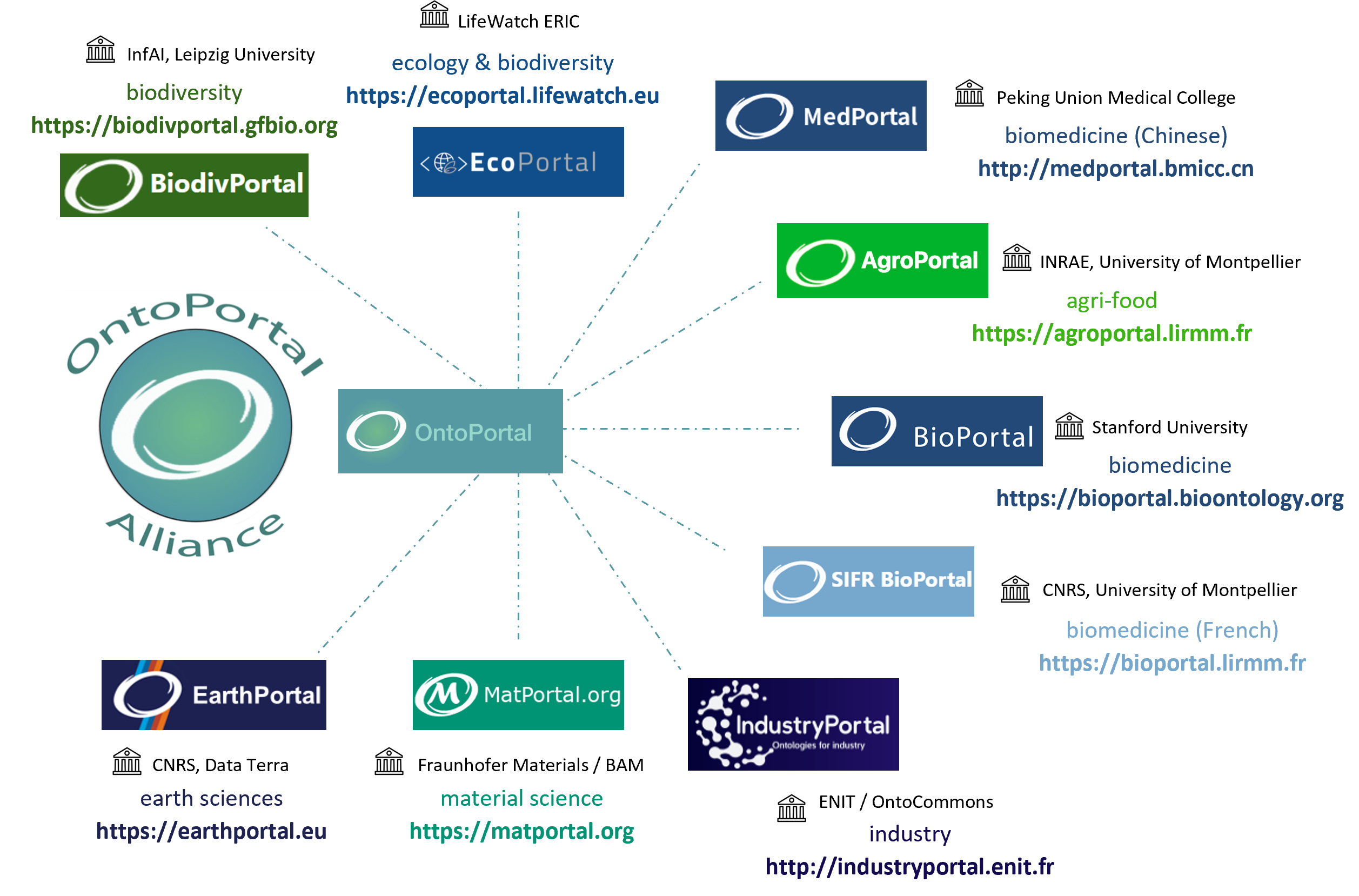
Teams in the Alliance develop and maintain several openly accessible ontology repositories and semantic artefact catalogues. These ontology repositories include BioPortal, the primary and historical source of OntoPortal code, but also AgroPortal, EcoPortal, MatPortal and more...
Many researchers use services of the OntoPortal virtual appliance in dozens of local installations with content and capabilities tailored to their own research needs. Several teams also use OntoPortal to host public semantic artefact catalogues and are banding together to advance development and community support for this key semantic tool.
Our motivation
The OntoPortal Alliance's original motivation and vision was to reuse outcomes and experiences obtained in the biomedical domain ––an area where the use of ontologies has always been important–– to serve and advance other scientific disciplines.
The OntoPortal Alliance has a main goal of synchronizing and sharing research and development efforts.
The group’s motivations are:
- to represent OntoPortal adopters and end users;
- to maximize the OntoPortal state-of-the-art service portfolio;
- to improve OntoPortal software while managing several parallel and different installations;
- to increase semantic uptake in science communities;
- to improve OntoPortal software while managing several parallel and different installations.
Our values
The Alliance is committed to be an open community.
By setting up a common repository on GitHub for the OntoPortal open source software, and by creating other shared resources for technical and strategic collaboration, the community can contribute to OntoPortal’s development, dissemination, and documentation. Together the Alliance plans the software’s improvements, adoption strategies, and long term growth and sustainability.
The code packaged and running within the appliance is available on the OntoPortal GitHub and licensed BSD-2, so every administrator or developer can easily get the relevant branches or forks and redeploy the code in the appliance. We strongly encourage Alliance partners (and other open-source contributors) to fork the OntoPortal GitHub repositories to enable traceability and collaborative contributions via their pull requests.
Besides code sharing, we use GitHub for issues, discussions, decision-making and overall project management.
Our roadmap
In 2022, we launched an annual 3-day-workshop that we see as key to fostering our growing community.
See the report of the 2022 edition.
The Alliance relies on the support provided by each project deploying an installation. We have implemented a free licensing system that we use to trace the many applications of the technology. In some cases, if the Alliance collaborates with industry or the private sector, we discuss the appropriate terms and conditions and possible financial participation.
Our community

Beyond the domain-specific portals, the OntoPortal technology is deployed by many external parties with other objectives. For instance, hospitals reuse the technology in-house to use services such as the Annotator on sensitive data.
Through 2015, the virtual appliance file was downloaded or deployed more than 140 times. Since version 3.0 (2020), 98 unique accounts have registered 135 OntoPortal appliances and in 2022, 60 unique appliance IDs called home including 19 running in Amazon Machine Instances.